Context of the study
The environmental DNA or eDNA method is a powerful tool for rapidly conducting exhaustive inventories of biodiversity in aquatic environments. However, applying it to establishing inventories fauna and flora on land is much more complex.
Indeed, the DNA present in a pond or a river is relatively homogeneously distributed thanks to the way the water mixes. In addition, it is relatively easy to filter large volumes in the field to concentrate the DNA before returning to the laboratory. In contrast, the physics of a terrestrial environment means that DNA left behind by mammals, birds or plants is relatively difficult to recover. Imagine having to carry dozens of kilos of soil to the laboratory!
At E-BIOM, we have been working for several years on establishing protocols to obtain the most exhaustive possible list of species present in a given environment by combining samples from different matrices.


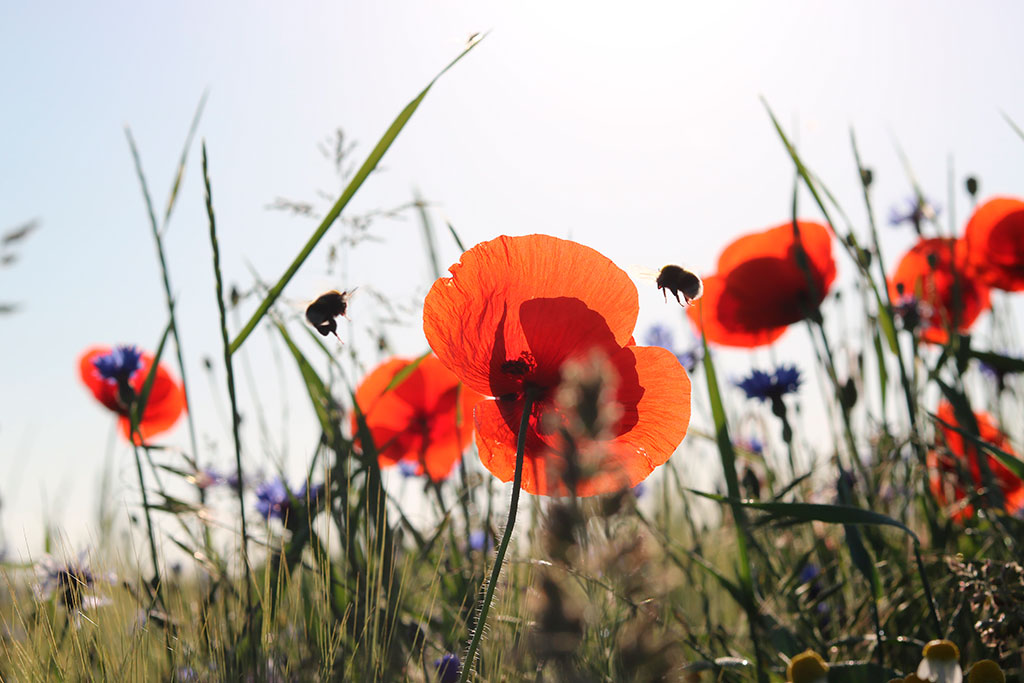
Working with beekeepers, naturalist associations and businesses, we have carried out a feasibility study to determine the value and limitations of the environmental DNA method when it comes to studying the floristic composition of honey samples produced by honey bees (Apis mellifera).
By gathering pollen from dozens, if not hundreds, of species of flowers, bees act as DNA collectors, concentrating it in balls of pollen and honey. This technique is particularly useful if you want to establish an inventory of melliferous plants in the vicinity of hives and so assess the ecological state of the surrounding environment, or if you are trying to determine the geographical regions of origin of different honeys.
Sampling and laboratory analysis
Honey samples were taken from a number of hives located in different environments: agricultural, urban, next to forests or near bodies of water… For some hives, the samples were repeated several times in order to study seasonal variations.
In parallel, botanical inventories were carried out by identifying the melliferous plants present in the area in which bees from different hives collected pollen, on the basis of morphological characteristics, in order to calibrate the method.
In the laboratory, the samples first had to be processed to separate the pollen grains contained in the honey from the sugar matrix. The DNA contained in the pollen grains was then extracted and amplified by PCR using universal primers targeting plant species. This method, known as environmental DNA metabarcoding, is particularly well suited to the study of complete taxonomic groups.
In order to guarantee the quality of our analyses and the reliability of our data, positive and negative controls were carried out to validate each step of the experimental process.


Results and conclusion
The results of the study are surprising! Up to 132 different species of melliferous plants were detected in a single sample of honey from honey bees. Environmental DNA analysis means that the floristic composition of honey samples can be studied, thus complementing palynology and the identification of pollen grains using microscopy.
Collecting honey samples at different times of the year has also shown seasonal variation in the way honey bees collect pollen and which plant species they visit. These results are consistent with traditional botanical surveys!
Lastly, the detection of indicator species means that we can infer the types of habitats located near the hives. These data give us an overview of the quality of the environment, as well as allowing us to determine the geographical areas from which the honey comes.
To go one step further, we then compare the results obtained from the honey samples with those generated by the analysis of pollen grains collected at the entrance of Apis mellifera hives or directly collected from honey bees and wild bees.